Introduction
REST Data Access Web Service Reference Specification
Page Contents
A summary of specific versions, resources, and XML namespaces is provided below.
- Supported Versions
- Changes in Current Revision
- Requesting an API Version
- REST
- API Data Types
- XML Data Elements
- XML Data Types
The REST Data Access Web Service is the fundamental interface in the GenoLogics Rapid Scripting API. Data is provided in XML with data access methods via HTTP.
*REST* (Representational State Transfer) is an architectural design style, not a specification. The term REST was incorporated into the API's name because the RESTful design style is perfect for research lab data access: self-descriptive, self-contained data representation in formats widely applicable to general scripting languages and the internet.
This REST reference section contains specifics and details on each resource, supported methods, and XML elements. For details on any resource click on the *REST resources* tree view at the left of this page. For details on the XML used by each resource method, use the *XML Namespaces* tree view.
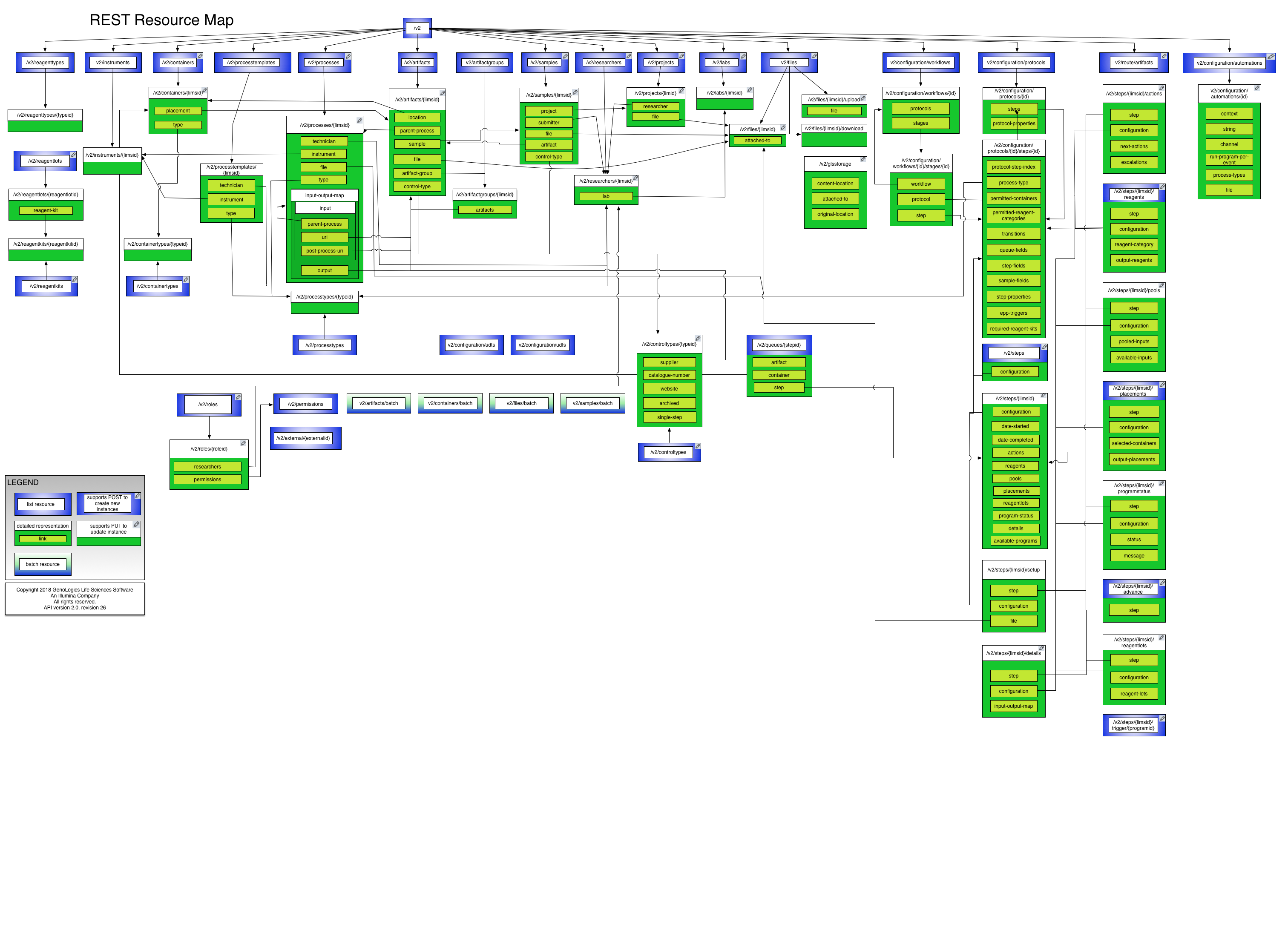
A good place to start is in the REST General Concepts section. To help navigate the resources, the following diagram highlights the methods available and the major resource relationships.
Supported Versions
API changes are tracked with the combination of a version number and a revision number.
The version number indicates forwards and backwards compatibility, while the revision number within the version describes features added to the API that will not negatively affect current functionality. Only the version number is referenced as part of the request; the revision number simply tracks incremental enhancements to the API.
The following version is described in this section of the web site:
| Version | Current Revision | Released with |
|---|---|---|
| v2 | r26 | Clarity LIMS 5.1 |
Previous versions can be accessed like so: http://genologics.com/developer/{ClarityVersion}
For example: http://genologics.com/developer/4.2
----Changes in Current Revision
- Demux endpoint has been added for individual artifacts: /api/v2/artifacts/{limsid}/demux
- Added LIST/GET endpoints for step automations:
- /api/v2/configuration/automations
- /api/v2/configuration/automations/{id}
- Added Pre-release POST/PUT endpoints for step automations:
- /api/v2/configuration/automations
- /api/v2/configuration/automations/{id}
- Automations can have files attached to them. The files endpoint will accept an automation uri as a valid attach-to entity.
- Removed ability to update the attach-to attribute of a file on PUT.
- Added 'original-name' element to the file resource.
- Added 'automationname' and 'originalname' parameters to the file LIST endpoint.
- PUT/POST on processtype with a process parameter (step automation) has changed behaviour:
- Will attempt to reuse existing identical automations before creating a new one.
- It will no longer be possible to change the details of an existing automation.
- Automation can now be specified via uri.
- We have deprecated the custom fields on process-type's process-output elements.
- These fields are no longer honored when running steps in Clarity or using the steps API – all of the output's custom fields can be written.
- However, the output's field associations will continue to be honored when using the processes API.
- Added DELETE to sample: /api/v2/samples/{limsid}
Requesting an API Version
The version number is a path parameter in each resource URI in the API. The desired API version is substituted into the following request URI:
/{version}/{resource}
The following example would request version 2 of the containers resource:
- /v2/containers
REST
This API supports a Representational State Transfer (REST) model for accessing a set of resources through a fixed set of operations. The following resources are accessible through the RESTful model:
- /{version}
- /{version}/{version}
- /{version}/artifactgroups
- /{version}/artifactgroups/{experimentid}
- /{version}/artifacts
- /{version}/artifacts/{limsid}
- /{version}/artifacts/{limsid}/demux
- /{version}/artifacts/batch/retrieve
- /{version}/artifacts/batch/update
- /{version}/configuration/automations
- /{version}/configuration/automations/{automationid}
- /{version}/configuration/protocols
- /{version}/configuration/protocols/{protocolid}
- /{version}/configuration/protocols/{protocolId}/steps/{stepId}
- /{version}/configuration/udfs
- /{version}/configuration/udfs/{udfid}
- /{version}/configuration/udts
- /{version}/configuration/udts/{udtid}
- /{version}/configuration/workflows
- /{version}/configuration/workflows/{workflowid}
- /{version}/configuration/workflows/{workflowid}/stages/{stageid}
- /{version}/containers
- /{version}/containers/{limsid}
- /{version}/containers/batch/create
- /{version}/containers/batch/retrieve
- /{version}/containers/batch/update
- /{version}/containertypes
- /{version}/containertypes/{containertypeid}
- /{version}/controltypes
- /{version}/controltypes/{controltypeid}
- /{version}/external/{externalid}
- /{version}/files
- /{version}/files/{limsid}
- /{version}/files/{limsid}/download
- /{version}/files/{limsid}/upload
- /{version}/files/batch/retrieve
- /{version}/files/batch/update
- /{version}/glsstorage
- /{version}/instruments
- /{version}/instruments/{instrumentid}
- /{version}/labs
- /{version}/labs/{labid}
- /{version}/permissions
- /{version}/permissions/{permissionId}
- /{version}/processes
- /{version}/processes/{limsid}
- /{version}/processtemplates
- /{version}/processtemplates/{limsid}
- /{version}/processtypes
- /{version}/processtypes/{processtypeid}
- /{version}/projects
- /{version}/projects/{limsid}
- /{version}/queues/{protocolStepId}
- /{version}/reagentkits
- /{version}/reagentkits/{reagentkitid}
- /{version}/reagentlots
- /{version}/reagentlots/{reagentlotid}
- /{version}/reagenttypes
- /{version}/reagenttypes/{reagenttypeid}
- /{version}/researchers
- /{version}/researchers/{researcherid}
- /{version}/roles
- /{version}/roles/{roleId}
- /{version}/route/artifacts
- /{version}/samples
- /{version}/samples/{limsid}
- /{version}/samples/batch/create
- /{version}/samples/batch/retrieve
- /{version}/samples/batch/update
- /{version}/steps
- /{version}/steps/{limsid}
- /{version}/steps/{limsid}/actions
- /{version}/steps/{limsid}/advance
- /{version}/steps/{limsid}/details
- /{version}/steps/{limsid}/placements
- /{version}/steps/{limsid}/pools
- /{version}/steps/{limsid}/programstatus
- /{version}/steps/{limsid}/reagentlots
- /{version}/steps/{limsid}/reagents
- /{version}/steps/{limsid}/setup
- /{version}/steps/{limsid}/trigger/{programid}
There is also a WADL document describing the REST API.
API Data Types
All endpoints act on a common set of data. The data can be represented in different data formats (i.e. MIME types), depending on the endpoint that consumes and/or produces the data. The data can described by XML Schema, which definitively describes the XML representation of the data, but is also useful for describing the other formats of the data, such as JSON.
This document will describe the data using terms based on XML Schema. Data can be grouped by namespace, with a schema document describing the elements and types of the namespace. Generally speaking, types define the structure of the data and elements are instances of a type. For example, elements are usually produced by (or consumed by) a REST endpoint, and the structure of each element is described by its type.
API Data Type Schemas
| id | namespace | schema file |
|---|---|---|
| ri | http://genologics.com/ri | ri.xsd |
| art | http://genologics.com/ri/artifact | artifact.xsd |
| artgr | http://genologics.com/ri/artifactgroup | artifactgroup.xsd |
| aut | http://genologics.com/ri/automation | aut.xsd |
| cnf | http://genologics.com/ri/configuration | configuration.xsd |
| con | http://genologics.com/ri/container | container.xsd |
| ctp | http://genologics.com/ri/containertype | containertype.xsd |
| ctrltp | http://genologics.com/ri/controltype | controltype.xsd |
| exc | http://genologics.com/ri/exception | exception.xsd |
| file | http://genologics.com/ri/file | file.xsd |
| inst | http://genologics.com/ri/instrument | instrument.xsd |
| itp | http://genologics.com/ri/instrumenttype | instrumenttype.xsd |
| lab | http://genologics.com/ri/lab | lab.xsd |
| perm | http://genologics.com/ri/permission | permission.xsd |
| prc | http://genologics.com/ri/process | process.xsd |
| prx | http://genologics.com/ri/processexecution | processexecution.xsd |
| ptm | http://genologics.com/ri/processtemplate | processtemplate.xsd |
| ptp | http://genologics.com/ri/processtype | processtype.xsd |
| prj | http://genologics.com/ri/project | project.xsd |
| prop | http://genologics.com/ri/property | property.xsd |
| protcnf | http://genologics.com/ri/protocolconfiguration | protcnf.xsd |
| que | http://genologics.com/ri/queue | queue.xsd |
| kit | http://genologics.com/ri/reagentkit | reagentkit.xsd |
| lot | http://genologics.com/ri/reagentlot | reagentlot.xsd |
| rtp | http://genologics.com/ri/reagenttype | reagenttype.xsd |
| res | http://genologics.com/ri/researcher | researcher.xsd |
| role | http://genologics.com/ri/role | role.xsd |
| rt | http://genologics.com/ri/routing | routing.xsd |
| smp | http://genologics.com/ri/sample | sample.xsd |
| stg | http://genologics.com/ri/stage | stage.xsd |
| stp | http://genologics.com/ri/step | step.xsd |
| protstepcnf | http://genologics.com/ri/stepconfiguration | protstepcnf.xsd |
| udf | http://genologics.com/ri/userdefined | userdefined.xsd |
| ver | http://genologics.com/ri/version | ver.xsd |
| wkfcnf | http://genologics.com/ri/workflowconfiguration | wkfcnf.xsd |
XML Data Elements
- namespace ri
- namespace art
- namespace artgr
- namespace aut
- namespace cnf
- namespace con
- namespace ctp
- namespace ctrltp
- namespace exc
- namespace file
- namespace inst
- namespace itp
- namespace lab
- namespace perm
- namespace prc
- namespace prx
- namespace ptm
- namespace ptp
- namespace prj
- namespace prop
- namespace protcnf
- namespace que
- namespace kit
- namespace lot
- namespace rtp
- namespace res
- namespace role
- namespace rt
- namespace smp
- namespace stg
- namespace stp
- namespace protstepcnf
- namespace ver
- namespace wkfcnf
XML Data Types
- namespace ri
- namespace art
- namespace artgr
- namespace aut
- namespace cnf
- namespace con
- namespace ctp
- namespace ctrltp
- namespace exc
- namespace file
- namespace inst
- namespace itp
- namespace lab
- namespace perm
- namespace prc
- namespace prx
- namespace ptm
-
namespace ptp
- container-type-link
- control-type-link
- epp-trigger
- field
- file
- icebucketfield
- invocation-type
- output-generation-type
- parameter
- process-input
- process-output
- process-type
- process-type-attribute
- process-type-link
- process-types
- queuefield
- reagent-kit-link
- step-property
- step-setup
- style
- trigger-point
- trigger-status
- trigger-type
- type-definition
- variability-type
- namespace prj
- namespace prop
- namespace protcnf
- namespace que
- namespace kit
- namespace lot
- namespace rtp
- namespace res
- namespace role
- namespace rt
- namespace smp
- namespace stg
-
namespace stp
- actions
- actions-link
- action-type
- artifact
- automatic-next-step-link
- available-program
- container
- creation-input
- details
- details-link
- escalated-artifact
- escalation
- escalation-request
- escalation-review
- file
- input
- input-output-map
- instrument
- next-action
- output
- output-generation-type
- output-placement
- placements
- placements-link
- pooled-inputs
- pools
- pools-link
- program-status
- program-status-link
- reagent-label
- reagent-lot-link
- reagent-lots-link
- reagents
- reagents-link
- reagents-lots
- setup
- setup-link
- status
- step
- step-configuration
- step-creation
- user
- namespace protstepcnf
- namespace udf
- namespace ver
- namespace wkfcnf